Scientific Highlights
-

Commercial Licensing of a BESC Global Genetic Regulator of Aromatic Biosynthesis
December 10, 2015. BESC has been studying SNPs and key compositional and conversion phenotypes in the natural variation population in Populus using GWAS (genome-wide association studies). A study of rare SNPs identified an unusual paralog in Populus which had taken on new regulator functions. Patent applications have been filed. Follow-on work is testing the utility in other plant species. The technology is being licensed to two companies who plan to commercialize the technology: GreenWood Resources and Forest Genetics International. Increasing biofuel yield or forage digestibility can lead to reductions in land use for these crops and thus contributing to long term sustainability.
Muchero, et al. "Transcription factor which regulates flavonoid, phenylpropanoid, tyrosine, and tryptophan pathways," provisional - USPTO 14/720,023. Filed December 10, 2015.
-
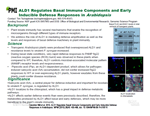
ALD1 Regulates Basal Immune Components and Early Inducible Defense Responses in Arabidopsis
April 2015. This paper addresses the role of ALD1 in mediating defense amplification as well as the levels and responses of basal defense machinery in plant immunity.
Cecchini NM et al. 2015. "ALD1 regulates basal immune components and early inducible defense responses in Arabidopsis," Molecular Plant-Microbe Interactions 28(4), 455–66. doi:10.1094/MPMI-06-14-0187-R.
-

Ten initial BESC Populus TOP lines identified with attractive biofuel traits in conversion and growth
February 21, 2015. Simultaneous comparison of Populus TOP lines for enhanced growth and biofuel traits will allow BESC researchers to identify the best among them to further evaluate in field studies (e.g., to determine sustainability traits) in preparation for application and licensing with commercial partners.
-

Toward improving tolerance of thermophilic microorganisms to pretreatment inhibitors
December 03, 2014. A heat-stable furfural reductase has been identified in T. pseudethanolicus 39E for inhibitor detoxification. It promises rational improvement in tolerance and has already shown promising results when expressed in the cellulolytic thermophile, C. bescii.
Clarkson et al. 2014. "A comparative multidimensional LC-MS proteomic analysis reveals mechanisms for furan aldehyde detoxification in Thermoanaerobacter pseudethanolicus 39E," Biotechnol. Biofuels 7(165). doi:10.1186/s13068-014-0165-z.
-

Transgenic American chestnuts show enhanced blight resistance and transmit the trait to T1 progeny
November 2014. The 'Darling4' transgenic is intermediate in resistance between susceptible American and resistant Chinese chestnut. Stem and leaf assays both show enhanced blight resistance due to the two transgenes (oxalic acid oxidase and ESF39, an antimicrobial peptide) in transgenic American chestnut trees. These genes have been transmitted to the T1 generation, which have shown similarly enhanced blight resistance. Metabolomic analyses indicate that concentrations are within the range observed for a panel of control plants and that the nuts of transgenic chestnut plants are likely safe for consumption.
Newhouse, Andy, et al. 2014. "Transgenic American chestnuts show enhanced blight resistanceand transmit the trait to T1 progeny," Plant Science 228, 88–97. doi:10.1016/j.plantsci.2014.04.004.
-

Overexpression of key cellulose biosynthesis precursor enzyme, UGPase, reveals altered sugar and secondary metabolism in Populus
October 07, 2014. This study provides evidence that manipulation of sugar metabolizing enzymes, such as UGPase, can have qualitative and quantitative effects on secondary metabolic pathways of shikimate, phenylpropanoid, and lignin biosynthesis. A cross-collaborative BESC effort is investigating the substrate preferences and reaction kinetics of UGPase to understand the observed pleiotropic phenotype. In integration with assessments of effects of other gene sequence and expression variations, our studies are yielding critical insights into biomass formation in Populus.
Payyavula RS, Tschaplinski TJ, Jawdy SS, Sykes RW, Tuskan GA and Kalluri UC. 2014. "Metabolic profiling reveals altered sugar and secondary metabolism in response to UGPase overexpression in Populus," BMC Plant Biology 14(265). doi:10.1186/s12870-014-0265-8.
-

New relationship between lignan biosynthesis and lignin distribution in secondary cell wall structure in Arabidopsis
August 05, 2014. Provides new evidence for a relationship between lignan synthesis and the role of lignin in secondary plant cell wall structure, which may shed light on evolution of plant secondary wall structure. Further analysis of this relationship may provide strategies for reducing recalcitrance of lignocellulosic biomass while enhancing plant defense.
Zhao, et al. 2014. "Pinoresinol reductase 1 impacts lignin distribution during secondary cell wall biosynthesis in Arabidopsis," Phytochemistry 112, 170-78. doi:10.1016/j.phytochem.2014.07.008.

Populus trichocarpa and Populus deltoides Exhibit Different Metabolomic Responses to Colonization by the Symbiotic Fungus Laccaria bicolor
June 2014. Shows that the metabolic changes during favorable colonization between the ectomycorrhizal fungus Laccaria bicolor and its compatible host, Populus trichocarpa, are characterized by shifts in aromatic acid, organic acid, and fatty acid metabolism. Demonstrates that this extensive metabolic reprogramming is repressed in incompatible interactions and that more defensive compounds are produced or retained. We also demonstrate that L. bicolor can metabolize a number of secreted defensive compounds and that the degradation of some of these compounds produces immune response metabolites (e.g., salicylic acid from salicin). Therefore, results suggest that the metabolic responsiveness of plant roots to L. bicolor is a determinant factor in fungus–host interactions.
Tschaplinski et al. 2014. "Populus trichocarpa and Populus deltoides exhibit different metabolomic responses to colonization by the symbiotic fungus Laccaria bicolor," Molecular Plant-Microbe Interactions 27(6), 546–56. doi:10.1094/MPMI-09-13-0286-R.
-

Lignin Valorization: Improving Lignin Processing in the Biorefinery is outlined in Science
May 15, 2014. This perspective review offers a rationale for optimism that more value can be derived from lignin within a biorefinery due to current and projected research.
Ragauskas, et al. 2014. "Lignin valorization: improving lignin processing in the biorefinery," Science 344(6185), 709–19. doi:10.1126/science.1246843.
-

Caldicellulosiruptor bescii (C. bescii) degrades lignin in switchgrass as well as cellulose and hemicellulose
May 01, 2013. Chemical and thermal pretreatment strategies for conventional cellulosic biofuels production contribute a significant cost overhead to the overall process. This unexpected result of simultaneous lignin and carbohydrate solubilization by C. bescii bodes well for complete industrial conversion by extremely thermophilic microbes of biomass that requires limited or no chemical pretreatment.
Kataeva, et al. 2013. "Carbohydrate and lignin are simultaneously solubilized from unpretreated switchgrass by microbial action at high temperature," Energy and Environmental Science 6(7), 2186–95. doi:10.1039/c3ee40932e.
-
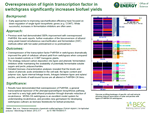

Overexpression of lignin transcription factor in switchgrass significantly increases biofuel yields
April 01, 2013. Results have demonstrated that overexpression of PvMYB4, a general transcriptional repressor of the phenylpropanoid/lignin biosynthesis pathway, can lead to a very high yield ethanol production through dramatic reduction of recalcitrance. MYB4-OX switchgrass is an excellent model system for understanding recalcitrance, and provides new germplasm for developing switchgrass cultivars as biomass feedstocks for biofuel production.
Shen, et al. 2013. "Enhanced characteristics of genetically modified switchgrass (Panicum virgatum L.) for high biofuel production," Biotechnology for Biofuels 6(71). doi:10.1186/1754-6834-6-71.
-
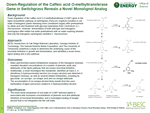

Down-Regulation of the Caffeic acid O-methyltransferase Gene in Switchgrass Reveals a Novel Monolignol Analog
September 28, 2012. The more facile breakdown of cell walls of COMT-deficient plants is associated with increased concentrations of phenolic acid and aldehyde inhibitors of microbial fermentation, and a monolignol analog of sinapyl alcohol that is not integrated into the cell walls.
Tschaplinski, et al. 2012. "Down-regulation of the caffeic acid O-methyltransferase gene in switchgrass reveals a novel monolignol analog," Biotechnology for Biofuels 5(71). doi:10.1186/1754-6834-5-71.
-

BESC Reports Clostridium thermocellum Ethanol Stress Responses
July 23, 2012. Largest responses were related to nitrogen uptake and metabolism Likely important for redirecting the cells physiology to overcome inhibition and allow growth to resume. Study suggests possible avenues for metabolic engineering and provides comprehensive, integrated systems biology datasets useful for future metabolic modeling, synthetic biology and strain development endeavors.
Yang, S. et al. "Clostridium thermocellum ATCC27405 transcriptomic, metabolomic and proteomic profiles after ethanol stress," BMC Genomics 13(336). doi:10.1186/1471-2164-13-336.
-

Pseudomonas fluorescens Induces Strain-Dependent and Strain-Independent Host Plant Responses in Defense Networks, Primary Metabolism, Photosynthesis, and Fitness
June 2012. Using Arabidopsis plants as a model, Populus deltoides bacterial isolate GM30 were found to be a plant-growth-promoting rhizobacterium (PGPR) increasing lateral root growth and protecting the plant host against disease. Root colonization of Arabidopsis by GM30 elicited a systemic defensive response that was elucidated by network modeling of gene expression and metabolite pathway analysis. This work identifies a specific gene network driving the systemic response in plant-PGPR interactions. This work also provides a baseline network for understanding the roles of multiple microbial associates and host plant genotypes.
Weston et al. 2012. "Pseudomonas fluorescens induces strain-dependent and strain-independent host plant responses in defense networks, primary metabolism, photosynthesis, and fitness," Molecular Plant-Microbe Interactions 25(6). doi:10.1094/MPMI -09-11-0253.