Biogeochemical Transformations at Critical Interfaces
ORNL Mercury Science Focus Area (SFA)
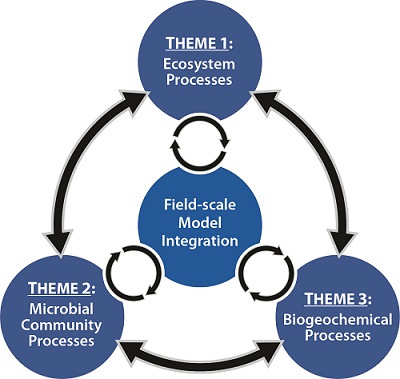
Theme 3: Biogeochemical Complexity and Molecular Mechanisms of Hg Transformations
The overarching goal of Theme 3 is to gain a fundamental understanding of complex biogeochemical processes and their interactions [e.g., dissolved organic matter (DOM), microbes, particulate organic matter (POM) and minerals, and water chemistry in EFPC] controlling Hg species transformation and availability for cellular uptake and methylation. Our specific objectives are to address the following scientific questions:
- What are the dominant Hg-binding organic ligands or molecular compositions (e.g., thiolates in DOM), and how do they competitively interact and control Hg speciation?
- What are the Hg-binding domains on cell membrane and cytosols, and how do cells competitively interact with extracellular organic and inorganic ligands for Hg binding, uptake (either passive or active), and ultimately methylation?
- How does environmental complexity (e.g., DOM, microbes, and minerals) influence Hg species distribution and availability for cell sorption, uptake, and methylation?
FY20–FY21 Accomplishments
Theme 3 made significant progress toward milestones over the past 12 months and has published 10 papers, with 10 additional manuscripts currently under review or in preparation. These studies are mostly focused on understanding a complex, yet finite set of geochemical and biomolecular processes controlling behavior, transformations, and net MeHg production in the environment. A robust predictive understanding of Hg biogeochemical transformations requires knowledge about the underlying molecular mechanisms and coupled interactions between Hg-binding organic and inorganic ligands, minerals, and methylating and demethylating microorganisms.
Overlooked Mercury Isotope Exchange in Environmental Tracer Studies and Implications
![Spontaneous Mercury (Hg) Isotope Exchange.
Illustration of spontaneous Hg isotope exchange
between enriched <sup>202</sup>Hg(0) spike and <sup>201</sup>Hg bound to various
environmental matrices, such as thiols and dissolved
organic matter (DOM), resulting in rapid redistributions
of Hg isotopes bound to the ligand. [Reprinted with permission
from Wang, Q., et al. 2020. “Rates and Dynamics
of Mercury Isotope Exchange between Dissolved Elemental
Hg(0) and Hg(II) Bound to Organic and Inorganic
Ligands.” Environmental Science & Technology Technology.
54(23):15534–45. DOI:10.1021/acs.est.0c06229. See
also Zhang, L., et al. 2021.] Spontaneous Mercury (Hg) Isotope Exchange](/programs/rsfa/images/2021/Fig8.jpg)
click to enlarge
Spontaneous Mercury (Hg) Isotope Exchange. Illustration of spontaneous Hg isotope exchange between enriched 202Hg(0) spike and 201Hg bound to various environmental matrices, such as thiols and dissolved organic matter (DOM), resulting in rapid redistributions of Hg isotopes bound to the ligand. [Reprinted with permission from Wang, Q., et al. 2020. “Rates and Dynamics of Mercury Isotope Exchange between Dissolved Elemental Hg(0) and Hg(II) Bound to Organic and Inorganic Ligands.” Environmental Science & Technology Technology 54(23):15534–45. DOI:10.1021/acs.est.0c06229. See also Zhang, L., et al. 2021.]
Enriched Hg-stable isotopes have been widely used as tracers in field and laboratory investigations of Hg biogeochemical transformations, such as methylation and demethylation. Few studies, however, have considered concurrent isotope exchange reactions between newly spiked and preexisting ambient Hg in environmental matrices. Using enriched Hg (as mercuric HgCl2), we examined isotope exchange between enriched Hg and ambient Hg(II) bound to various environmental matrices, including soil minerals, low-molecular-weight (LMW) thiols, and DOM. The impact of isotope exchange on MeHg production in the presence of organic ligands was evaluated with an iron-reducing bacterium, Geobacter sulfurreducens PCA. Surprisingly, we found that the spiked Hg(II) rapidly exchanged with ligand- or mineral-bound ambient Hg(II), resulting in redistribution of Hg isotopes bound to the ligands or minerals. We also observed an apparently similar methylation rate and magnitude of the spiked Hg(II) and ambient Hg(II) by PCA cells. These observations underscore the importance of isotope exchange when an enriched Hg isotope is applied in environmental matrices, as the exchange could potentially lead to biased rate calculations of Hg transformation and bioaccumulation and thus result in biased risk assessments of new Hg input to the natural ecosystems.
Additionally, we investigated the rates and dynamics of isotope exchange between dissolved elemental 202Hg(0)aq and 201Hg(II) bound to organic and inorganic ligands with varying chemical structures and binding affinities. Time-dependent exchange reactions were followed by isotope compositional changes using both inductively coupled plasma mass spectrometry and Zeeman cold vapor atomic absorption spectrometry. Rapid, spontaneous isotope exchange (<1 h) was also observed between 202Hg(0)aq and 201Hg(II)-bound to chloride (Cl–), ethylenediaminetetraacetate (EDTA), and thiols, such as cysteine (CYS), glutathione (GSH), and 2,3-dimercaptopropanesulfonic acid (DMPS). Without external reductants or oxidants, the exchange resulted in transfers of two electrons and redistribution of Hg isotopes bound to the ligand but no net changes of chemical species in the system. However, an increase in the thiol:Hg(II) ratio decreased the exchange rates due to the formation of 2:1 or higher thiol:Hg(II) chelated complexes, but it had no effects on exchange rates with 201Hg(II) bound to EDTA or Cl–. The exchange between 202Hg(0)aq and 201Hg(II) bound to DOM showed an initially rapid exchange rate followed by a slower exchange rate, likely resulting from Hg(II) complexation with both low- and high-affinity binding functional groups on DOM (e.g., carboxylates vs bidentate thiolates). These results demonstrate that Hg(0)aq readily exchanges with Hg(II) bound to various ligands and highlight the importance of considering exchange reactions in experimental enriched Hg isotope tracer studies or in studies where natural abundance Hg isotope are present in environmental matrices.
Roles of Methanobactin in the Biogeochemical Cycling of Hg and Transition Metals
Methanotropic bacteria and the chalkophore methanobactin have been implicated in the biogeochemical cycling of Hg. Methanobactins facilitate the acquisition of Cu(II) ions in some methanotrophic bacteria, analogous to siderophores and iron. Methanobactins are also known to form strong complexes with other late transition metals, including Hg. Thus, methanobactins influence the bioavailability of Hg(II) for microbial methylation (Yin et al. 2020) and MeHg for degradation by methanotrophs (Lu et al. 2017). The influence of methanobactins on MeHg production by anaerobic bacteria is considerable. Recent studies revealed significant differences between methanobactins produced by different methanotrophs, e.g. Methylosinus trichosporium OB3b and Methylocystis sp. strain SB2. We hypothesize that the chemistry and configuration of functional groups with the methanobactin metal binding site controls Hg(II) complexation, ligand exchange reactions involved in Hg(II) uptake by methylators, and ultimately methylation of Hg(II) by HgcAB.
To better understand the interplay between solution-phase configurations and metal interactions of methanobactins, we compared spectroscopic signatures of methanobactin with Hg(II) and several transition metal complexes. We studied the complexation of Zn, Cd, and Hg ions by methanobactin from Methylocystis sp. strain SB2 using a combination of absorbance, fluorescence, and extended X-ray absorption fine structure (EXAFS) spectroscopy, complemented by time-dependent density functional theory (TD-DFT) calculations. Characteristic changes in absorbance and fluorescence spectra occur over a wide range of experimental timescales and arise from a stoichiometric complexation with thiolate functional groups in methanobactin. Hg L3-edge EXAFS and TD-DFT calculations suggest a linear model for Hg-S coordination, while TD-DFT calculations are consistent with a tetrahedral model for Zn(II) and Cd(II). Enhancement of the fluorescence emission of methanobactin observed upon interaction with transition metals indicates a mechanism of complexation-hindered isomerization of the intrinsic fluorophores in methanobactin. Investigating the biomolecular basis of metal complexation advances our understanding of ligand exchange reactions controlling Hg(II) uptake and MeHg production.
Isolation of Two Methanotrophs from EFPC and Impact on Methanotrophic-Mediated MeHg Degradation
In collaboration with Prof. Jeremy Semrau at University of Michigan and Theme 1, two novel methanotrophs of the Gammaproteobacteria class, Methylomonas sp. EFPC1 and Methylococcus sp. EFPC2, were isolated from the Hg-contaminated EFPC biofilm samples. The 16S rRNA sequence analyses of Methylomonas sp. strain EFPC1 indicated that it was phylogenetically similar to Methylomonas sp. LW13, and Methylococcus sp. strain EFPC2 was most similar to Methylococcus geothermalis IM1T. Average nucleotide identity (ANI) values between Methylomonas sp. strain EFPC1 and Methylomonas sp. LW13 and between Methylococcus sp. strain EFPC2 and Methylococcus sp. IM1T were ~95% and 73%, respectively. Genes for particulate methane monooxygenase (pMMO) were found in both Methylomonas sp. strain EFPC1 and Methylococcus sp. strain EFPC2, while evidence of a divergent form of pMMO (pXMO) and soluble methane monooxygenase was found in only Methylomonas sp. strain EFPC1. Genes for the central pathway of methane oxidation were found in both methanotrophs, as well as for the ribulose monophosphate pathway and tricarboxylic acid cycle (Kang-Yun et al. 2021a).
![Methanobactin SB2-Mercury (Hg) X-ray Absorption Spectroscopy. (A) A comparison of the non-phase shift corrected Fourier transforms of the Hg L3-edge EXAFS data for 1:1 (black) and 2:1 (red) mixtures of mb-SB2:Hg2+. The inset shows the corresponding EXAFS comparison. (B) and (C) show the best structural fits (derived using FEFF) to the data for 1:1 and 2:1 mixtures of mb-SB2:Hg2+, respectively. [Reprinted from Eckert P., et al. 2021. “Spectroscopic and computational investigations of organometallic complexation of group 12 transition metals by methanobactins from Methylocystis sp.
SB2.” Journal of Inorganic Biochemistry. In press. DOI:10.1016/j.jinorgbio.2021.111496. Copyright 2021, with permission
from Elsevier.] Methanobactin SB2-Mercury (Hg) X-ray Absorption Spectroscopy](/programs/rsfa/images/2021/Fig9.jpg)
click to enlarge
Methanobactin SB2-Mercury (Hg) X-ray Absorption Spectroscopy. (A) A comparison of the non-phase shift corrected Fourier transforms of the Hg L3-edge EXAFS data for 1:1 (black) and 2:1 (red) mixtures of mb-SB2:Hg2+. The inset shows the corresponding EXAFS comparison. (B) and (C) show the best structural fits (derived using FEFF) to the data for 1:1 and 2:1 mixtures of mb-SB2:Hg2+, respectively. [Reprinted from Eckert P., et al. 2021. “Spectroscopic and computational investigations of organometallic complexation of group 12 transition metals by methanobactins from Methylocystis sp. SB2.” Journal of Inorganic Biochemistry. In press. DOI:10.1016/j.jinorgbio.2021.111496. Copyright 2021, with permission from Elsevier.]
Additionally, we found evidence for methanobactin “theft” among methanotrophs, which could potentially impact on methanotrophic-mediated MeHg degradation (Kang-Yun et al. 2021b). Aerobic methanotrophy is strongly controlled by copper (Cu), and methanotrophs are known to have multiple mechanisms for Cu uptake. Some methanotrophs secrete a MB that binds Cu with extremely high affinity, while others utilize a surfacebound protein (MopE) and a secreted form (MopE*) for Cu collection. Cu competition may therefore significantly impact methanotrophic community composition and activity. We show that Methylomicrobium album BG8, Methylocystis sp. strain Rockwell, and Methylococcus capsulatus Bath, all lacking genes for MB biosynthesis, are not limited for Cu by multiple forms of MB. Interestingly, Mm. album BG8 and Methylocystis sp. strain Rockwell were found to have genes similar to mbnT that encodes for a TonB-dependent transporter required for MB uptake, indicating that these methanotrophs may “steal” Cu-MB complexes and degrade MeHg. Indeed, substantial demethylation was observed by these strains in the presence of MB. Mc. Capsulatus Bath, however, lacks anything similar to mbnT, and it was unable to degrade MeHg either in the presence or absence of MB. Similarly, when mbnT was deleted in Mm. album BG8, MeHg degradation in the presence of MB was indistinguishable from when MB was not added. These results indicate that methanotrophic-mediated MeHg degradation may be more widespread than previously thought and thus play an important role in net production of MeHg in the environment.
 ORNL Mercury SFA sponsored by Subsurface Biogeochemical Research (SBR) program, U.S. Department of Energy's Office of Biological and Environmental Research. Paul Bayer, SBR Program Manager.
ORNL Mercury SFA sponsored by Subsurface Biogeochemical Research (SBR) program, U.S. Department of Energy's Office of Biological and Environmental Research. Paul Bayer, SBR Program Manager.
Security Notice | Contact Eric Pierce, ORNL | Website Questions | Site Map
File last modified: Wednesday, September 01, 2021
The ORNL Mercury SFA is sponsored by the Subsurface Biogeochemical Research (SBR) program within the U.S. Department of Energy's Office of Biological and Environmental Research.