Biogeochemical Transformations at Critical Interfaces
ORNL Mercury Science Focus Area (SFA)
Featured Research Highlight
Kinetics of Enzymatic Mercury Methylation at Nanomolar Concentrations Catalyzed by HgcAB
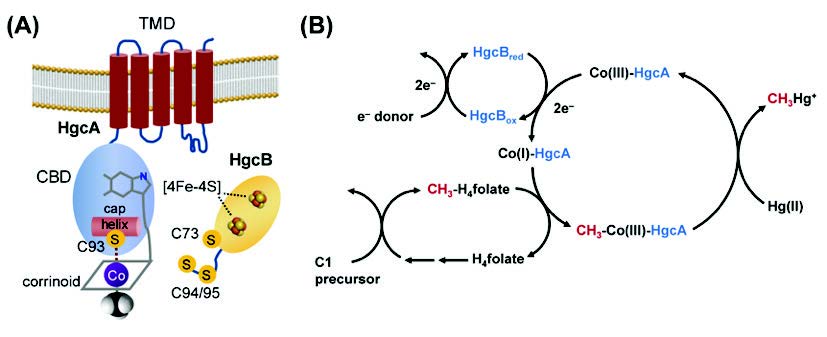
Structural features of HgcA and HgcB and proposed mechanism of HgcAB-mediated Hg methylation. (A) Schematic representation of HgcA and HgcB. TMD = transmembrane domain, CBD = corrinoid-binding domain, [4Fe-4S] = iron-sulfur cluster, C = cysteine. (B) Proposed roles of the proteins HgcA and HgcB in Hg methylation. C1 = one-carbon precursor, Co(I)/(III) = cobalt center of corrinoid cofactor and its oxidation state, H4folate = tetrahydrofolate. CH3H-4folate = 5-methyltetrahydrofolate, Hg(II) = mercuric mercury, CH3Hg+ = methylmercury.
Study reveals insights into the biochemistry of mercury methylation by anaerobic bacteria expressing the gene pair hgcAB
The Science
Methylmercury (MeHg) is a potent bioaccumulative neurotoxin, which is produced by certain anaerobic bacteria and archaea. Mercury (Hg) methylation has been linked to the gene pair hgcAB encoding a membrane-associated corrinoid protein and a ferredoxin. Although microbial Hg methylation has been characterized in vivo, the cellular biochemistry and the specific roles of the gene products HgcA and HgcB in Hg methylation are not well understood. Here we report the kinetics of Hg methylation in cell lysates of Desulfovibrio desulfuricans ND132 at nanomolar Hg concentrations. The enzymatic Hg methylation mediated by HgcAB is highly oxygen-sensitive, irreversible, and follows Michaelis-Menten kinetics with an apparent KM of 3.2 nM and Vmax of 19.7 fmol·min–1·mg–1 total protein for the substrate Hg(II). Although the abundance of HgcAB in the cell lysates is extremely low, Hg(II) was quantitatively converted to MeHg at sub-nanomolar substrate concentrations. Increasing thiol/Hg(II) ratios did not impact Hg methylation rates, which suggests that HgcAB-mediated Hg methylation effectively competes with cellular thiols for Hg(II) consistent with the low apparent KM. Supplementation of 5-methyltetrahydrofolate or pyruvate did not enhance MeHg production, while both ATP and a non-hydrolyzable ATP analog decreased Hg methylation.
The Impact
Exposure to neurotoxic MeHg through the consumption of fish represents a significant risk to human health. Anaerobic microbial communities in sediments and periphyton biofilms have been identified as sources of MeHg in aquatic systems, but the underlying biomolecular mechanisms are not fully understood. This study provides insights into the biochemistry of Hg methylation in sulfate-reducing bacteria.
Summary
Anaerobic bacteria that carry hgcAB generate MeHg in aquatic environments. Gaining insights into the biochemistry of Hg methylation is important for understanding factors driving Hg methylation. HgcA and HgcB catalyze the formation of MeHg at extremely low Hg(II) concentrations and effectively compete with cellular thiols. Advancing our understanding of microbial MeHg production may inform strategies to curtail the formation of neurotoxic MeHg in the environment.
Publication
Date, S. S., J. M. Parks, K. W. Rush, J. D. Wall, S. W. Ragsdale, and A. Johs. 2019. “Kinetics of enzymatic mercury methylation at nanomolar concentrations catalyzed by HgcAB.” Applied Environmental Microbiology. DOI: 10.1128/AEM.00438-19.
 ORNL Mercury SFA sponsored by Subsurface Biogeochemical Research (SBR) program, U.S. Department of Energy's Office of Biological and Environmental Research. Paul Bayer, SBR Program Manager.
ORNL Mercury SFA sponsored by Subsurface Biogeochemical Research (SBR) program, U.S. Department of Energy's Office of Biological and Environmental Research. Paul Bayer, SBR Program Manager.
Security Notice | Contact Eric Pierce, ORNL | Website Questions | Site Map
File last modified: Thursday, August 08, 2019
The ORNL Mercury SFA is sponsored by the Subsurface Biogeochemical Research (SBR) program within the U.S. Department of Energy's Office of Biological and Environmental Research.