Biogeochemical Transformations at Critical Interfaces
ORNL Mercury Science Focus Area (SFA)
Featured Research Highlight
Key Amino Acid Residues for Mercury Methylation Confirmed
Team constructs site-directed mutants of HgcA and HgcB proteins and examines the effect on production of methylmercury pollutant.
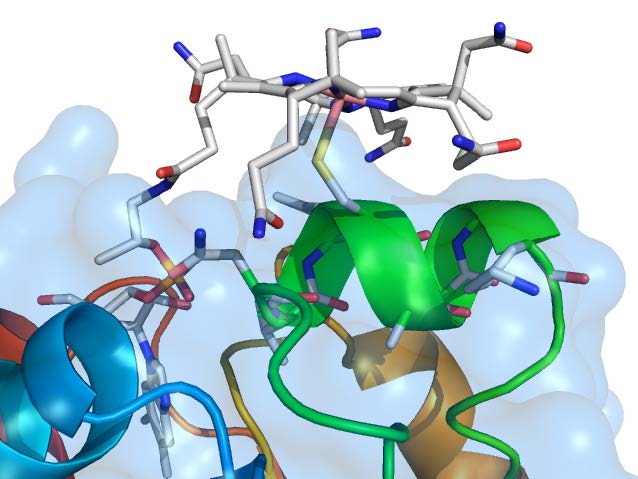
Homology model of the corrinoid-binding domain of HgcA from Desulfovibrio desulfuricans ND132. The corrinoid cofactor is illustrated at the top of the figure (cobalt shown in pink). The proximity of the strictly conserved Cys (sulfur in yellow) to cobalt in the corrinoid suggests a unique "base-off" "Cys-on" configuration and thus a possible functional role for this interaction in the methylation reaction. The "cap-helix" is shown as a green ribbon (N90–K97). Amino acid side chains are represented as stick models. Additional elements are oxygen (red), nitrogen (blue), and carbon (gray).
The Science
Methylmercury is a potent neurotoxin produced from inorganic mercury by anaerobic microbes in natural environments worldwide. Once produced, methylmercury accumulates in the aquatic food chain, thus posing a risk to human health in many regions. Recently, the two genes necessary for microbial production of methylmercury were identified: hgcA encoding a corrinoid protein and hgcB encoding a ferredoxin. To date, all microbes possessing orthologs of these genes have been found to be capable of methylating mercury, whereas organisms lacking these genes are not. Although these findings firmly establish the significance of these proteins in mercury methylation, mechanistic details about their predicted roles in the process require further study. In this research, the proteins were explored to determine whether amino acid residues predicted to be important in methylmercury formation could be mutated and how these mutations affect methylmercury production.
The Impact
This research reveals new insight into mercury methylation by anaerobic microorganisms. Results support the previously predicted importance of an amino acid residue (Cys93) in HgcA and reveal additional residues in both proteins that facilitate methylmercury production. In addition, these structural details may help identify alternative physiological functions of these proteins that could point to evolutionary drivers for maintaining the capacity of anaerobes to methylate mercury.
Summary
The deduced amino acid sequence of HgcA was threaded onto the crystal structure of the corrinoid iron-sulfur protein (CFeSP), revealing a cysteine (Cys93) to be within liganding distance of the cobalt of the corrinoid of HgcA. This finding supported the predicted transfer of a carbanion or methyl radical from the corrinoid to the mercuric ion, leading to methylmercury formation. Mutation of Cys93 to Ala93 or Thr93 completely eliminated the methylation capacity, whereas His93 retained a small methylation capacity. Thus, the previously predicted importance of the strictly conserved Cys93 in HgcA was confirmed, as was the "cap helix" orienting the Cys93 to the cobalt (see figure). Surprisingly, HgcB could not be compensated by other ferredoxins apparently encoded in the genome.
Publication
Smith, S. D., R. Bridou, A. Johs, J. M. Parks, D. A. Elias, R. A. Hurt Jr., S. D. Brown, M. Podar, and J. D. Wall. 2015. "Site-directed mutagenesis of HgcA and HgcB reveals amino acid residues important for mercury methylation." Appl. Environ. Microbiol. 81:3205–17. [DOI:10.1128/AEM.00217-15]
 ORNL Mercury SFA sponsored by Subsurface Biogeochemical Research (SBR) program, U.S. Department of Energy's Office of Biological and Environmental Research. Paul Bayer, SBR Program Manager.
ORNL Mercury SFA sponsored by Subsurface Biogeochemical Research (SBR) program, U.S. Department of Energy's Office of Biological and Environmental Research. Paul Bayer, SBR Program Manager.
Security Notice | Contact Eric Pierce, ORNL | Website Questions | Site Map
File last modified: Tuesday, February 04, 2020
The ORNL Mercury SFA is sponsored by the Subsurface Biogeochemical Research (SBR) program within the U.S. Department of Energy's Office of Biological and Environmental Research.