Biogeochemical Transformations at Critical Interfaces
ORNL Mercury Science Focus Area (SFA)
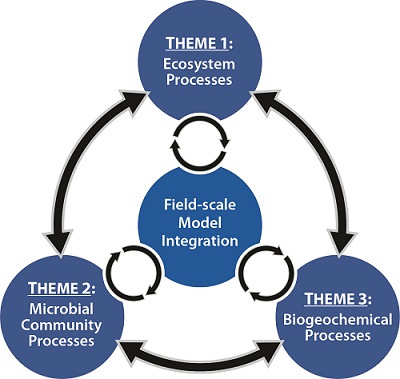
Theme 2: Microbial Community Processes
The overall goals of Theme 2 are to (1) understand the mechanisms of Hg methylation at the molecular scale and the consequences to the cell in planktonic and biofilm lifestyles, whether in isolation, synthetic, or natural microbial communities; (2) determine the breadth and depth of Hg-methylating species; and (3) elucidate the biochemical roles of HgcA and HgcB. Our research is designed to answer the following questions:
- How widespread is the ability to methylate Hg, and what are the relative contributions from different microbial clades to the overall net pool of MeHg generated in different types of environments, specifically in EFPC?
- What genes and metabolic traits are required for function and maintenance of hgcAB?
- What environmental conditions alter HgcAB expression?
- What is the biochemical (native) function of HgcA and HgcB in the absence of Hg?
- Can sequence-inferred HgcAB structural models provide a mechanistic framework for testing structure-function hypotheses of Hg binding, methylation, and potential involvement of other proteins in the methylation process?
- Do mutations to hgcAB affecting Hg methylation also change organismal fitness under certain environmental conditions?
- Does the overall cellular metabolism and MeHg generation change in multispecies cultures versus single-organism cultures?
FY20–FY21 Accomplishments
Over the past 12 months, Theme 2 has published a number of manuscripts and made significant progress toward our milestones, including (1) determining alternative (native) functions of HgcAB, (2) identifying and attempting to isolate novel Hg methylators from EFPC sediments, (3) developing model microbial communities to reflect the EFPC community, (4) determining the role sulfide mineral surfaces have in mediating dimethylmercury (DMeHg) formation.
click to enlarge
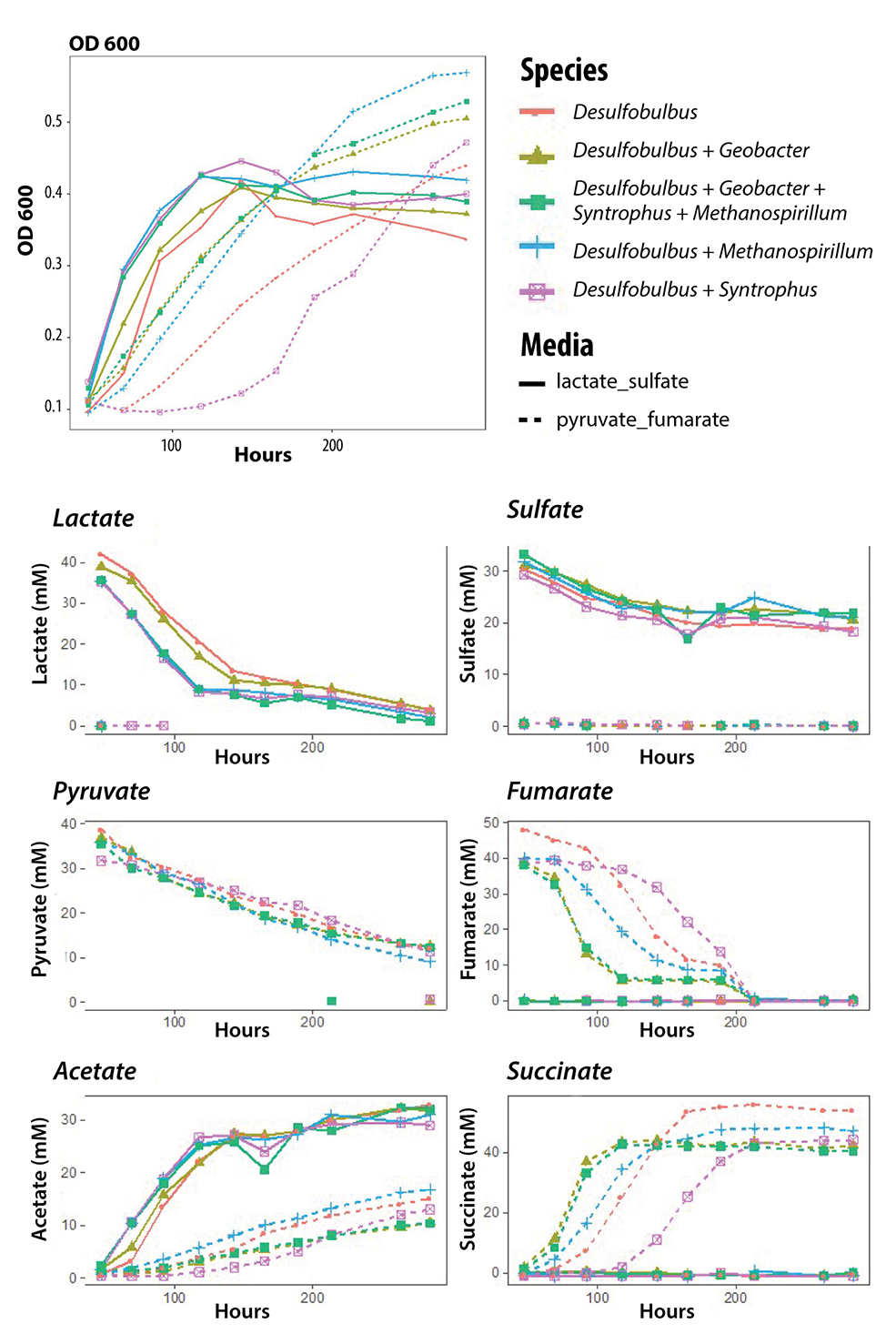
Assessing hgcAB Alternative Function. Left: Growth curves of the individual and various multicultures used in this study. Right: The temporal loss of electron donors and acceptors, lactate/sulfate or pyruvate/fumarate, and the production the donor and acceptor degradation products, acetate and succinate, respectively. Solid lines represent lactate/sulfate cultures and broken lines represent pyruvate/fumarate cultures.
Determining Alternative hgcAB Functions
We have continued work related to physiological experiments for determining the native biochemical function of HgcAB. In FY21, we completed numerous batch culture bottle experiments with Desulfovibrio desulfuricans ND132 wild-type and mutant strains grown in defined media with various substrates (e.g., pyruvate, fumurate, lactate, sulfate, formate, and acetate). We chose mutant strains related to carbon and Hg cycling that exhibited differences in Hg-methylation capability compared to the wild-type (e.g., 0 to 246%). Additionally, we continue to analyze large omic datasets obtained from the Environmental Molecular Sciences Laboratory (EMSL; user proposal 50174). Results showed differences in substrate consumption, acetate production, and transcription of one-carbon (C1) metabolism genes between mutant strains and wild-type under fermentative and sulfate-reducing conditions. However, results were not consistently within a given pathway but rather seemed to be random. Most notably, mutants deficient in methylation also did not express genes for motility, including cilia and flagella. Reasons for the lack of motility gene expression are not immediately clear.
We have hypothesized that the native physiological function of HgcAB may be related to C1 metabolism for acetyl-CoA and methionine biosynthesis, metal resistance, or metalloid methylation. Clues to the native biochemical function of HgcAB may lie in determining the environmental conditions that control expression and translation of hgcA. Therefore, for the last year, we have been investigating transcriptional regulation of hgcA under different growth parameters using D. desulfuricans ND132 as a model organism. We utilized both molecular reverse transcription–quantitative polymerase chain reaction (RT-qPCR) and metaomic (RNA-seq) methods to test whether changes in hgcA expression occurred when wild-type ND132 cells were grown in conditions that require the postulated biochemical functions of HgcAB (e.g., +/– formate, methionine, arsenate, and Hg). Indeed, our results show that hgcA expression is significantly regulated across the growth stages of D. desulfuricans ND132 under some conditions tested. Specifically, we found that there is an arsR-like gene upstream of hgcAB and have hypothesized that, as in As metabolism, this gene may control hgcAB expression. To date, we have tested hgcAB expression with different As forms and precursors for methionine production. These tests have revealed that both hgcAB expression and Hg methylation were altered in several of the conditions, including with arsenate and arsenite.
![Dimethylmercury Formation on Cadmium Sulfide Surfaces. Calculated reaction pathway for DMeHg formation on a model CdS nanoparticle. Charges on selected atoms are labeled. [Reprinted with permission from Lian, P., et al. 2021. Mechanistic investigation of dimethylmercury formation mediated by a sulfide mineral surface. Journal of Physical Chemistry.
DOI: 10.1021/acs.jpca.1c04014. Copyright 2021 American Chemical Society.] Assessing hgcAB Alternative Function](/programs/rsfa/images/2021/Fig7.jpg)
click to enlarge
Dimethylmercury Formation on Cadmium Sulfide Surfaces. Calculated reaction pathway for DMeHg formation on a model CdS nanoparticle. Charges on selected atoms are labeled. [Reprinted with permission from Lian, P., et al. 2021. "Mechanistic investigation of dimethylmercury formation mediated by a sulfide mineral surface." Journal of Physical Chemistry. DOI: 10.1021/acs.jpca.1c04014. Copyright 2021 American Chemical Society.]
Developing Model Microbial Communities and Fitness Assay
In FY21, we have made steady progress toward understanding the effect of multispecies interactions on MeHg generation and developing model microbial communities to reflect the EFPC community. Large-volume (1- to 4-L), single-species and dual-species batch cultures have been completed in bottles and bioreactors to characterize growth and chemical profiles for methylating and nonmethylating sulfate-reducing bacteria and methanogens, including Desulfobulbus propionicus, Desulfobulbus oligotrophicus, Methanospirillum hungatei, and Methanococcus maripaludis. We measured a suite of analytical parameters throughout the growth curve for each culture, including growth rates based on optical density (OD) and protein content, Hg methylation, MeHg demethylation, hgcaA expression, qPCR and fluorescence in situ hybridization (FISH) determination of relative cell abundance, simple metabolites (anions/cations/organic acids), sulfide concentration, and pH (see Fig. 6). Methylation/demethylation analyses are ongoing due to the pandemic, but the pairings of methylating and nonmethylating organisms allow us to disentangle the specific effect of syntrophic interactions on MeHg generation in organisms with very different metabolisms (i.e., sulfate reduction versus methanogenesis). To date, all culturing and analysis, with the exception of the Hg analyses, are complete. Continuing efforts are underway in FY21 to increase the complexity of multispecies cultures to recapitulate the range of metabolisms present in EFPC sediment, including the addition of Syntrophus aciditrophicus and G. sulfurreducens.
With respect to cell fitness under laboratory conditions, deletion of hgcAB does not significantly change the growth rate of D. desulfuricans ND132. However, maintenance of these genes across evolutionary time scales suggests that they provide an important fitness benefit in the environment. We hypothesize that the fitness effects of an hgcAB deletion are environmentally dependent and that these genes are conditionally dispensable under standard laboratory conditions. In conditions where hgcAB are providing their native function, we expect to see a fitness defect in a ΔhgcAB strain compared to the wild-type. So far, we have measured competitive fitness between wild-type and ΔhgcAB ND132 in more than 20 different media compositions. Although additional experiments are needed, our current results show that the strains grow differently when mixed together compared to when grown separately, despite minimal genetic differences between the two strains. In turn, this result suggests that HgcAB is involved in interactions between cells of ND132. Ongoing work is exploring these interactions in more detail.
Dimethylmercury Formation Mediated by a Sulfide Mineral Surface
Previous experimental work has shown that abiotic formation of dimethylmercury (DMeHg) from methylmercury (MeHg) can be enhanced on sulfide mineral surfaces. In collaboration with Niri Govind at Pacific Northwest National Laboratory (PNNL) and Sofi Jonsson at Stockholm University, we led a computational study that explored the mechanisms of DMeHg formation on the surface of a hawleyite nanoparticle. We showed that coordination of MeHg substituents to adjacent reduced sulfur groups protruding from the surface facilitate DMeHg formation and that the reaction proceeds through direct transmethylation from one MeHg substituent to another (see Fig. 7). Coordination of Hg by multiple S atoms provides transition state stabilization and activates a C-Hg bond for methyl transfer. Calculated energetics are in good agreement with available experimental data, providing confidence in the models. These findings fill a knowledge gap in our understanding of environmental Hg cycling.
 ORNL Mercury SFA sponsored by Subsurface Biogeochemical Research (SBR) program, U.S. Department of Energy's Office of Biological and Environmental Research. Paul Bayer, SBR Program Manager.
ORNL Mercury SFA sponsored by Subsurface Biogeochemical Research (SBR) program, U.S. Department of Energy's Office of Biological and Environmental Research. Paul Bayer, SBR Program Manager.
Security Notice | Contact Eric Pierce, ORNL | Website Questions | Site Map
File last modified: Wednesday, September 15, 2021
The ORNL Mercury SFA is sponsored by the Subsurface Biogeochemical Research (SBR) program within the U.S. Department of Energy's Office of Biological and Environmental Research.